Wow! it’s the sixth week, seems like time never stops and always keeps moving. Well today is a Monday, which are never a good feeling but we gotta hang in there. I started my day by going over the results with Professor Andresen, in order to come out with a solution or a new procedure. Eventually professor Andresen did came out with a new procedure which is the same procedure has before but with slightly different trial digestion. Instead of creating different samples of our solution by adding different amounts of micrococcal nuclease, we instead only made two 50 micro-liters of solution which contain different amounts of micrococcal nuclease. I places this solutions into the heater for 37 degrees Celsius and took out 5 micro-liters of each solution at a specific time. The different times that we assigned were 5, 20, and 40 minutes, with another sample that lasted about 2 hours. This procedure accomplished the task of creating different sizes of DNA by letting the microccocal nuclease eat up the nucleic acids and taking small amounts of sample at different times and stopping the process by adding EDTA. Once this process was finish, I continued the trial digestion with the same procedure as before by using proteinase K and SDS. In the mean while, I created a new gel in order to put the trials in tomorrow. I also made more TBE buffer to used the next day. All this procedure took approximately the whole day due to the unfortunately broken wrist I got. All is well and I could still work, and that’s really important to me because I never give up on something I start.
Final day of Week #5. Finally able to see Results
Its Friday! It is always good to come to work on a Friday because you feel at easy with yourself and work just for the fact that once this day is over, you can rest for two whole days.
Well I started my day out by making more TBE buffer, which consist of 50 ml of TBE (Tris-Borate-EDTA buffer) and 450 ml of DI water. Once I was done making this buffer I made two DNA ladder, DNA ladder is a solution of DNA molecules of different known lengths and is used to run along side our samples in order to estimate the size of our samples. Once this was done, I prepared an extra micro-centrifuge tube with only 1 micro-liter of our chromatic supernatant, 9 micro-liters of 60% sucrose and 10 micro-liters of DI water. I made this extra sample in order to see how it looks on the gel without the trial digestion. For the solutions who already had already been through the trial digestion, I took 1 micro-liter from each and added them to new micro-centrifuge test tubes with the same numbers label in order to not get mix up. I also added 9 micro-liters of 60% sucrose and 10 micro-liters of DI water to the new test tubes. Once this was done I was ready to add them into my gel. The solutions on the gel can be seen in figure #1.
 |
| Figure #1 Samples added to Gel |
Once I loaded all of my samples into the gel, I set the voltage to 100 volts and waiting until a blue line reached 5 cm. While I was waiting for this process to happen, I started to unpack all of the materials that we ordered in order to have more stock available to used in the future. It was like Christmas morning, unpacking all of the boxes. See figure #2
 |
| Figure # 2 Material needed for present and future usage |
Once the unpacking was done, our shelves finally looked like they had material in it and it looks awesome. See figure #3
 |
| Figure #3 More beakers to make more Buffer |
Week Five
The results from the last experiment I mentioned in my last blog post were inconclusive. Along with the next few experiments that had occurred after that. Now I’m not saying that it is impossible to get the data that we wanted to get with the materials that I was using, however, Professor Andresen pointed out that if the experiment was this finicky then no one would care about it enough to try to reproduce it. So later that day he found me a brand new batch of nano particles. This time we got Citrate coated gold nano particles and this changed our procedure slightly. The general idea now is to take the negatively charged gold nano particles and wrap positively charged PAH around the gold. Then we’ll wrap the DNA around the PAH/Gold complex. I was very excited to get away from the old procedure that I couldn’t figure out how to fix. The first problem we ran into with this new experiment was trying to coat the gold with the PAH. When I followed the old procedure something strange happened.
5th week, final steps on the preparation of Mononucleosomes
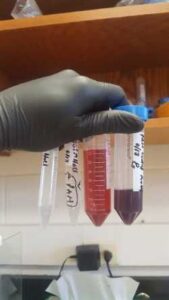
When I came back on Wednesday, I took out my solutions from the refrigerator and stop the spinner. This was a struggle but not because it was hard removing the equipment from the refrigerator but for removing it with a wrist paint I felt during this process. This was due to the accident I had with my skate board but as a hard worker I still manage to complete my task and keep working. Once the solution was out of the refrigerator, I put the solution into a special test tube which you are able to see in figure #1 and you are also able to see my wrist support that one of my lovely friend let me borrow.
 |
| Figure #1 Special test tube for Centrifuge, this is test tube is used when using a powerful Centrifuge that goes faster than 3900xg |
I set the centrifuge to a speed of 5000xg for 5 minutes and after this was done, I removed the supernatant and measured the Concentration of DNA using the UV-vis equipment. From our collected data, professor Andresen and myself found our that we lost about 300 mg of chromatin from our previous UV-vis sample but that is to expected when doing this kind of washing and processes. Once this was done, I dialyses my supernatant against 10mM Tris-HCI and 1mM CaC12 at 4 degrees Celsius. I was only able to create three dialysis bags for my supernatant due to the fact that my supernatant volume was a lot, the rest of the supernatant will be placed in dialysis bags the next day.
The next day I came back, I removed the dialysis bags from the buffer and created the rest of the dialysis bags for the rest of my supernatant. Once this was done I started the process for the Trial Digestion of my mononucleosomes. I prepared six different microcentrifuge tubes and label them with different numbers, shown in figure #2.
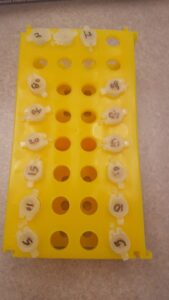 |
| Figure #2 Two rows of different samples and different concentrations of Micrococcal Nuclease |
Figure #2 shows two different rows of solutions, one solution was made up of 50 micro-liters of our supernatant and different concentration of Micrococcal nuclease. Once again Micrococcal nuclease is used to digest nucleic acids. We set this solutions to heat up for 55 minutes at a temperature of 37 degrees Celsius and once this is done we removed them from the heater and add 0.5 micro-liters of EDTA to stop this reaction. When we are done with all of this, we again prepare six other micro-centrifuges tubes with the same numbers but with an extra PK on the side to represent Proteinase K. This new tubes contain five micro-liters of our digested chromatic solution, five micro-liters of proteinase K, and 0.5 micro-liters of SDS. We then set this samples into the heater for 50 minutes at 50 degrees Celsius. Once this is done, we can placed our samples in the refrigerator to be used for the next day. During the time I was waiting for my solutions to heat up, I was making new Agarose gel in order to used it the next day and placed my samples in it.
 |
| Figure #3 Never give up! even if you feel like your hand is going to fall. |
5th week, Continuation of the preparation of Mononucleosomes
I started this new week by re-suspending my nuclei in 8ml of ML and spind it down at 3000xg for 5 minutes. I did this process 4 times in total and all of this is done for the preparation of Mononucleosomes. Once this was done four times, I removed the supernatant and took a small sample in order to check for how much DNA concentration I had and added 2M of CaC12 to make 1mm. Once this was done I need to make a calculation of how much Micrococcal nuclease to add to each sample. Micrococcal nuclease is used to eat up the DNA strands until it reaches the nucleus. Once I found the correct amount of Micrococcal nuclease to add, I pre-equilibrated the solution to 37 degrees Celsius for short DNA fragments for 30 minutes. Once the timer is up, in order to stop the reaction, I must add 10mm of EDTA, for my specific amount of solution, I added 161.488 micro-liters of EDTA. When all of this process was done, I was once spun the solutions at 1000xg for 5 minutes at 4 degrees Celsius. The final I step I needed to do was to create dialysis bags and let them sit over night, In my case I made 4 different dialysis bags, 3 containing our supernatant and the other containing the nuclei (left over white solution). The next pictures show the tools used to make the dialysis bags.
 |
| Dialysis tubing and Clips. |
 |
| Dialysis bag |
 |
| Dialysis bags, all inside 500mL of EDTA. This must be left over night. |
When 30-day free trial comes to your rescue
NITPIC can be used to extract the raw ITC data from NanoAnalyze to perform a number of functions including thermogram reconstruction and processing integrated isotherms from the raw thermogram. I ran a test trial using data from my 5mM DNA-Cobalt Hexammine data. The program succeeded in producing a beautiful isotherm for me. I then converted this isotherm into a format that is readable by SEDPHAT. SEDPHAT processed this data and subsequently produced a global fit. However, I still couldn’t figure out how to integrate this data with isotherms from my other trials, in order to generate an average fit for my data points. I played around with the program for hours to figure something out. But this is one of those stories without a happy ending.
 |
| NITPIC |
 |
| SEDPHAT |
However, when there’s a will, there’s a way. I discovered another way to generate a mean isothermal plot. This process was lengthy and relatively complex.
 |
| Sigmoid fit for the 5mM DNA isotherm. (And yes, the demo prints are due to the trial version) |
Week 3 in a nutshell
I started week 3 by performing an ITC run with the same 5 mM DNA-NaCl solution from last week to check for experimental inconsistencies. The results displayed on the screen after 1.5 hours of continuous ITC humming produced well-consistent data. In order to stretch the plots between the binding and condensation phases even further, I created an 8 mM DNA-NaCl sample and ran it by the ITC machine. The integrated heat peak for the binding phase had been successfully stretched further after this ITC run, resulting in a larger number of plots between the phases. The experimental parameters were very similar in the two runs that I performed using the 8mM DNA-NaCl solution.
Furthermore, I also performed a second heat of dilution for the 6mM Cobalt Hexammine solution in order to account for one anomalous point in the first run. Subsequently, I subtracted this data from the other DNA-Cobalt Hexammine reactions in order to standardize the binding enthalpies involved in those reactions. This was week 3 inside the lab in a nutshell.
Outside the lab, we had an amazing time at the Tuesday brown-bag lunch. Everybody involved with the X-Sig summer research program on campus took part and we had our fair share of laughter and discussion there. On Wednesday, we had a dinner meet at the Quarry Pavilion. The food was amazing! I had a lot of fun outside work during this week.
The rest of week # 4 Preparation for new sample of chicken blood
The rest of the week I kept washing the chicken blood with KTM and Triton X-100 in order to wash away the fat surrounding the cell. In order to wash the blood, I used 20 micro-liters of KTM and 70 micro-liters of Triton X-100 for each sample and spin it in the centrifuge with a force of 3600xg for 10 minutes at 4 degrees Celsius. Throughout the day I just kept washing the chicken blood and made more KTM because I was running out. This process kept on going from Thursday to Friday because I still had red solution in my sample which is not what I was looking for. This process continued until I can obtain white/clean nuclei. On Friday I was able to see results from washing the chicken blood and I could see white/clean nuclei for two of my four samples. I ran the same procedure for the two solutions once again and I was finally able to get white/clean nuclei for all my solutions. Figure #1 shows how the chicken blood must look after all the washes with KTM and Triton X-100.
 |
| Figure #1 White clean Nuclei |
After this was done, I once again wash the clean nuclei with just KTM in order to get rid of the Triton X-100. I did this procedure twice and by the time I was done the day was over. The washing of the chicken blood takes a good amount of time due to the amount of Triton X-100 that I added but for future reference I would recommend starting with at least 80 to 100 micro-liters of Triton X-100 and go down on the volume from there.
First Days of week #4 Trial and Error
I started out my week by once again doing the Trial Digestion chromatin due to the results we kept on getting from our DNA and Gel test. I started my work where I create samples with proteinase K, which contain 5 micro-liters of digested chromatin that I had previously created, 5 micro-liters of Proteinase K that professor Andresen recently bought, 0.5 micro-liters of SDS and for this new trial I added 0.55 micro-liters of CaC12 to the sample. After all this solutions where added, I let the samples sit for 50 minutes at a temperature of 50 degrees Celsius. While I was waiting for this to be done, I started preparing the gel once again. Once the gel was done, I placed the gel on the refrigerator for about 1 hour and 30 minutes. Figure one depicts the machine use to heat up the samples.
 |
| Figure #1 Isotemperature |
Once the 50 minutes were up, I took out the samples and placed them into ice. By this time I got new micro-centrifuge tubes and took 1 micro-liter of solutions from samples and added it to the new test tubes. I also added 9 micro-liters of 60% sucrose and 10 micro-liters of DI water. I did this for each sample that we had, and I also added 1 micro-liter of blue dye in order to see it once we added into the gel. During this time I also made DNA latter which contain about 4 micro-liter of DNA ladder and 10 micro-liter of 60% sucrose. During this time I took out the gel out of the refrigerator but notice that I used the wrong Comb! and this was really bad because due to this I needed to redo the gel and this took about the whole day. Once I was done making the gel I let it sit on the refrigerator though out the night. In my spare time I kept washing the new chicken blood with KTM. Figure # 2 displays the right comb to used for the gel.
 |
| Figure # 2 Comb used to make holes in the gel. |
The next day I came back, I took out the gel out of the refrigerator once again and this time I had everything ready to go so I put my samples in the gel and used a machine to provide voltage in order to move the DNA from one side to another. This process took about 5 hours and in the mean time I kept washing the Chicken blood with KTM. Once I came back from lunch, my gel was ready to be view by a UV machine. My results showed a better improvement on the last trials but It still wasn’t good enough for what we were looking for. Now professor Andresen and I decided to redo everything from the beginning but this time we will used the new fresh blood that was bought recently and see how this results come out and compare them our previous results. We aren’t so sure why our results came out the way they came out but we are thinking it might have been with storing our chicken blood in the -80 degrees Celsius refrigerator might have something to do with it. I guess this is part of being a researcher, to keep trying and improving on previous results in order to progress and grow both has a researcher but also has a human being.
Week #3 Results from our chromosomes
 |
| Figure 1# Machine used to heat up solutions |
 |
| Figure #2 Blue line where it needs to stop once DNA moves to the right due to charges. |
 |
| Figure #3 Shows the line where you must fill gel with TBE buffer |
 |
| Figure #4 Results from UV light and our chromosomes |