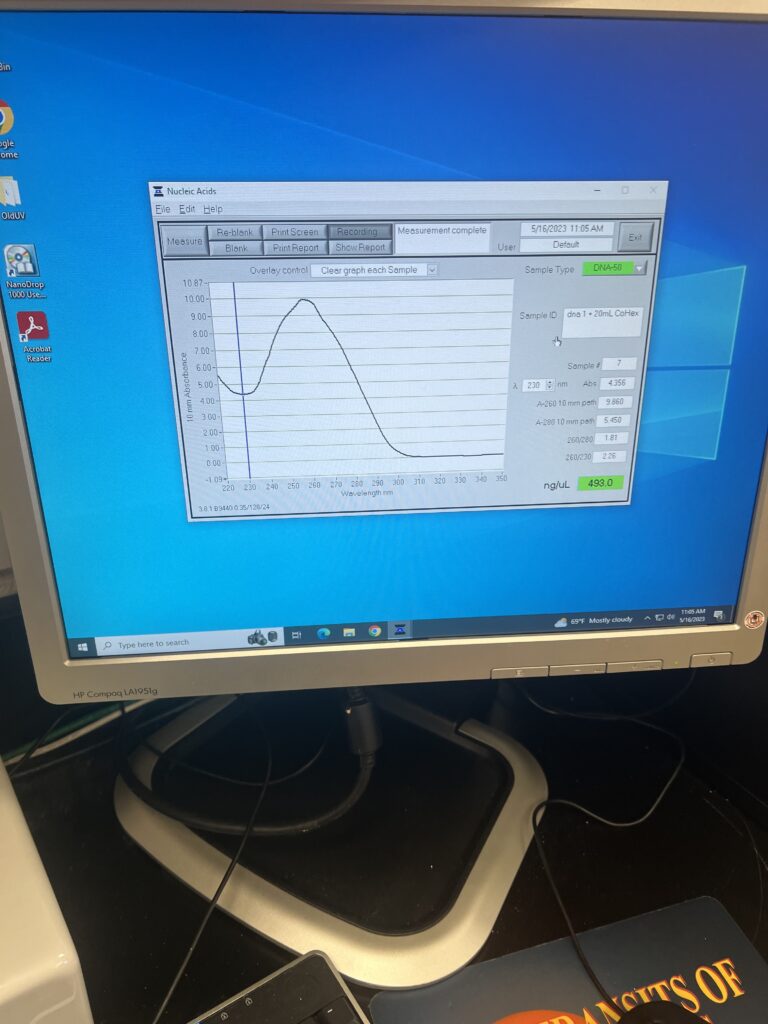
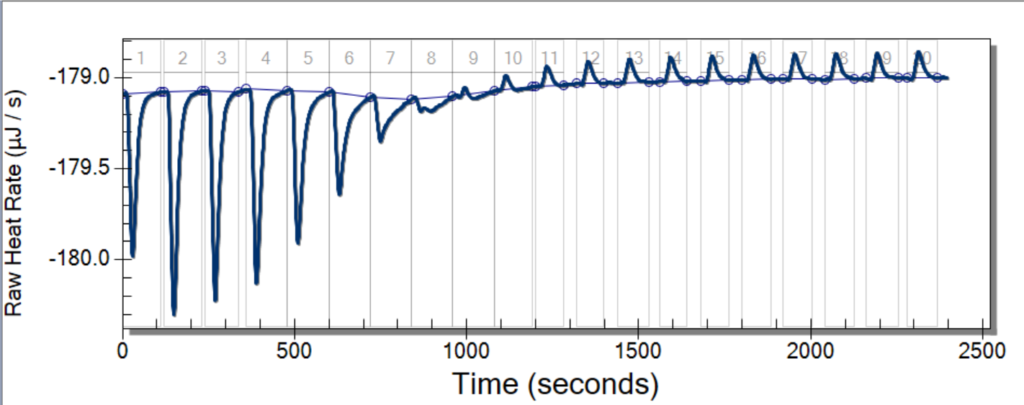
To start off this new week of research, we had a great team meeting to talk about our progress and, also, to discuss our goals for the week. For my partner, Aubrie, and me, our goal was to properly run the ITC. The Isothermal Titration Calorimetry is a process that involves keeping the equilibrium between a reference, which is usually water, and another sample that is being injected by another substance. The ITC then measures how the injections change the temperature of the reference and decide what it must take to keep the system (again, between the sample and reference) in equilibrium.
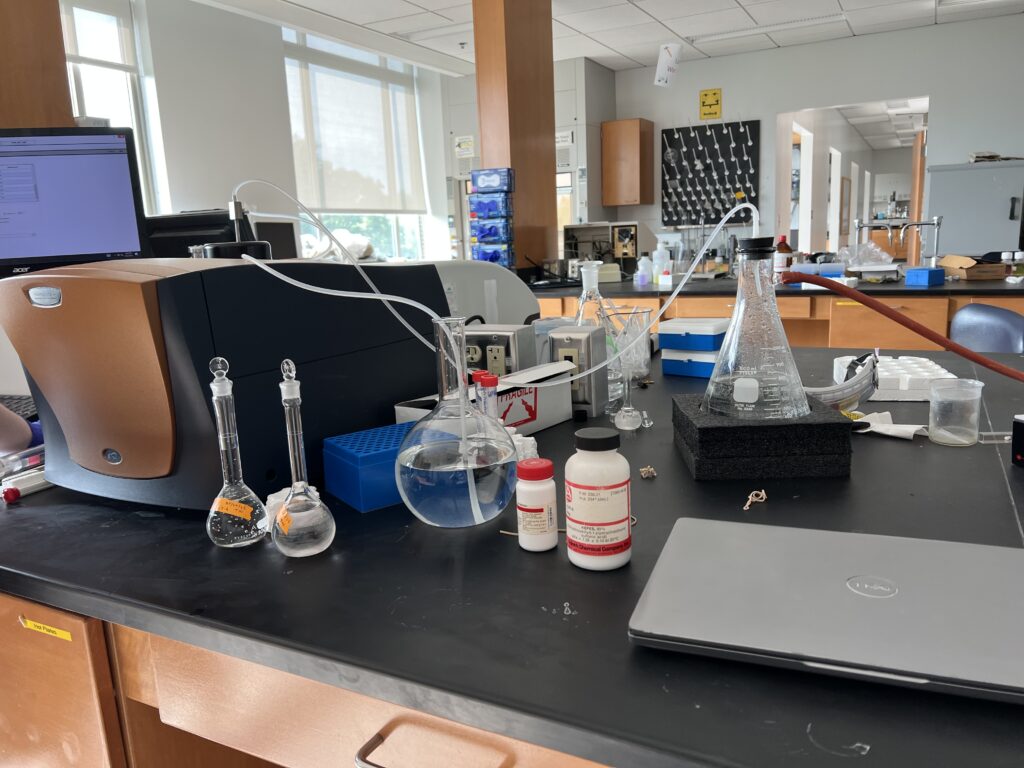

To train on the ITC, we tried to replicate the results of an experiment from a chemistry class taught by Professor Thompson. In this experiment, we injected EDTA, an acid, with magnesium. After a doing a total of ten runs of the experiment, we were able to get good data from four of those. The ITC is a particularly temperamental piece of equipment, which is why it can be so hard to get good data.
Once we decided that we were trained enough to run the ITC, we began our actual project. For this, we would need to make some DNA solutions with a Tris-EDTA buffer. Fun fact: our DNA actually comes from salmon testes! Although, we are having a bit of trouble getting some new DNA (perhaps due to salmon shortages) after our last few orders were cancelled.
Since having a TE buffer in the ITC can cause problems, we needed to wash this DNA with the buffer that we would actually be using in the ITC; sodium chloride. To do this, we dispersed a bit of our original solution into smaller tubes and combined it with even parts of a NaCl solution we made. We spun this in our centrifuge, who is now named Spinch, for a total of three times before we were able to wash the DNA completely with the sodium chloride. Warning: there were more steps in this process, please do not try to replicate at home.
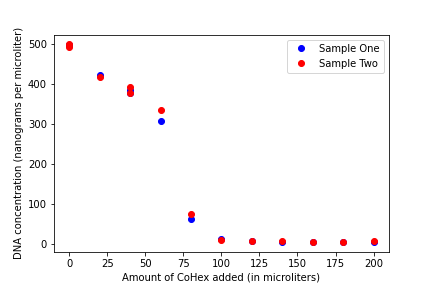
Another necessary ingredient for the experiment was a Cobalt Hexamine solution in sodium chloride. This turned out to be a very easy process and, before we knew it, we were ready to head over to the ITC and begin our very first run for our project!
For this project, we will be using DNA as the reference and the CoHex for the injections. We hope to replicate the data from last summer’s project as a starting point. We left the experiment running overnight and hope to see good results in the morning! For the last day of the week, we hope to run more of these experiments and try to problem-solve any issues that we may run into.
Until I have to clock in again,
Macyn.